Review of Species Interactions Available Through ChecklistBank
On Wed 22 October 2025, Rod Page gave a talk [1] in Bogotá, Columbia mentioning the “Afromoths” project by De Prins & De Prins [2] as an example. Afromoths is said to cover “all relevant information on every Afrotropical moth species” of Sub-Saharan Africa and includes descriptions of over 12k associated (plant) hosts (e.g., Daphnis nerii has host Nerium oleander L.) and over 500 parasites records (e.g., Agrionympha capensis has parasite Mesocomys pulchriceps Cameron) [3].
Facilitated by the Belgian Biodiversity Platform, Belspo, the Afromoth data is currently available through ChecklistBank in the Catalogue of Life Data Package (ColDP) format ([4], https://github.com/CatalogueOfLife/coldp). The ColDP format, developed by Markus Döring, Geoff Ower and colleagues, includes explicit support for species interactions, similar to how Darwin Core supports species interactions via their Resource Relationship Extension.
An example of the first record in a 2026 copy of their ColDP archive with sha256 and md5 fingerprints [5] hash://sha256/dd778e9038f87067815f9c7afdec1286db2e4cc08a298cbab27fe23dba2e1b44 and hash://md5/7e21ac93fb0e5e786f1254d6bd0a2341 respectively, was generated using the following bash script along with Preston v0.11.6 and miller:
preston cat \
--algo md5 \
--anchor hash://md5/a9aaa7113716ab2f87ede6f6b70297c8 \
--remote https://zenodo.org \
'line:zip:hash://md5/7e21ac93fb0e5e786f1254d6bd0a2341!/speciesinteractions.csv!/L1,L2' \
| mlr --icsv --oxtab cat
producing:
taxonID S100010010
relatedTaxonID
relatedTaxonScientificName Mesocomys pulchriceps Cameron
type has parasite
referenceID 6968
remarks
This record claims some reference with id 6968 that a taxon with id S100010010 has a parasite with a scientific name Mesocomys pulchriceps Cameron .
After extending Elton’s support to include Catalogue of Life Data Package, you can now access the record without having to join this records with their taxonomic and reference tables via an interpreted interaction record obtained via a published GloBI data review resource indexed-interactions.tsv.gz [3]:
preston cat \
--remote https://zenodo.org \
hash://md5/0c2373f08dc68ec2a44d18edb45d7139 \
| gunzip \
| mlr --itsvlite --oxtab \
filter '$sourceTaxonId == "S100010010" && $targetTaxonName == "Mesocomys pulchriceps Cameron"'
or, by replaying the associated archived review process [2] and generating interpreted interaction records from the archived ChecklistBank dataset using Elton v0.16.10 via
preston ls \
--algo md5 \
--anchor hash://md5/a9aaa7113716ab2f87ede6f6b70297c8 \
--remote https://zenodo.org \
| elton stream \
--algo md5 \
--data-dir data \
--prov-dir data \
--anchor hash://md5/a9aaa7113716ab2f87ede6f6b70297c8 \
--remote https://zenodo.org \
| mlr --itsvlite --oxtab \
filter '$sourceTaxonId == "S100010010" && $targetTaxonName == "Mesocomys pulchriceps Cameron"'
both of which are expected to produce a result including:
argumentTypeId https://en.wiktionary.org/wiki/support
[...]
sourceTaxonId S100010010
sourceTaxonName Agrionympha capensis
[...]
interactionTypeId http://purl.obolibrary.org/obo/RO_0002445
interactionTypeName hasParasite
[...]
targetTaxonName Mesocomys pulchriceps Cameron
[...]
referenceCitation Prinsloo G. L. & Uys V. M. (Eds.), 2015, Insects of cultivated plants and natural pastures in Southern Africa. vol issue pages i–xiv, 1–785
namespace urn:lsid:checklistbank.org:dataset:2017
archiveURI https://api.checklistbank.org/dataset/2017/archive.zip
lastSeenAt 2026-03-31T15:23:43.269Z
contentHash dd778e9038f87067815f9c7afdec1286db2e4cc08a298cbab27fe23dba2e1b44
eltonVersion 0.16.10
Discovery and Independent Review of ChecklistBank Associated Interaction Data
To help facilitate automated discovery of species interaction claims embedded in ChecklistBank Datasets, Elton, a GloBI commandline tool, is now able to list datasets with interaction claims using:
elton ls --online --registry checklistbank
On 2026-04-20, running the Elton command above produced the following list of datasets:
urn:lsid:checklistbank.org:dataset:1032
urn:lsid:checklistbank.org:dataset:1049
urn:lsid:checklistbank.org:dataset:1061
urn:lsid:checklistbank.org:dataset:1133
urn:lsid:checklistbank.org:dataset:1166
urn:lsid:checklistbank.org:dataset:1199
urn:lsid:checklistbank.org:dataset:2017
urn:lsid:checklistbank.org:dataset:2169
urn:lsid:checklistbank.org:dataset:2207
urn:lsid:checklistbank.org:dataset:2317
urn:lsid:checklistbank.org:dataset:2362
urn:lsid:checklistbank.org:dataset:124661
urn:lsid:checklistbank.org:dataset:265709
This suggests that a little over 10 datasets in ChecklistBank currently describe at least one species interaction claim. As expected, the Afromoth dataset (i.e., [2], urn:lsid:checklistbank.org:dataset:2017) is included in this list. Other examples include the Universal Chalcidoidea Database (i.e., [6], urn:lsid:checklistbank.org:dataset:124661) and the Catalogue of the Pterophoroidea of the World (i.e., [7], urn:lsid:checklistbank.org:dataset:1199).
This list of dataset identifiers can then be used to track associated datasets, and, if desired, generate interpreted species interaction records. For instance, for the Afromoths dataset with id urn:lsid:checklistbank.org:dataset:2017, a tab-seperated values table of their interpreted interaction data can be generated via:
elton track 'urn:lsid:checklistbank.org:dataset:2017'
elton interactions 'urn:lsid:checklistbank.org:dataset:2017' \
> interactions.tsv
assuming that ChecklistBank Web API is reachable, the tool elton is available, and the Afromoths dataset has the CoLDP format. These assumptions are expected to no longer hold in the near future because websites come and go, software tool versions may no longer be available, and dataset formats change or become obsolete.
To have a more resilient approach to capturing the tracked dataset and the interpretation process, the associated resource can be captured in a so-called “Bill of Material” which includes the digital fingerprints (or signatures) of the resources needed for Elton to interpret the dataset. This approach is used in the GloBI review process, and can be summarized as follows:
elton track --algo md5 \
--prov-mode \
'urn:lsid:checklistbank.org:dataset:2017' \
| elton tee --algo md5 \
| preston append --algo md5 \
| elton stream --algo md5 --data-dir data --prov-dir data \
| head -n2
in which the first line tells elton to generate the Data Bill of Materials (DataBOM) to document the versioned resources used to track, and interpret, the data associated with urn:lsid:checklistbank.org:dataset:2017 in rdf/nquads via the --prov-mode argument. Following, elton stores the associated resource in the data folders and the DataBOM is appended to a local Preston archive. In the second to last line (i.e., elton stream ...), elton is instructed to generate interaction records associated this DataBOM. Finally, in the last line, the first two lines in the resulting interaction record table are printed, resulting, at time of writing 2026-04-21, in the following record expressed in XTAB format:
The results in the data/ folder now contains all resources needed to reproduce this result, including the Data Bill of Materials outlining the process and versioned resources used. Because this DataBOM is a text file expressed in rdf/nquads, their digital fingerprint uniquely identifies the process and associated resources. Now, this DataBOM fingerprint (or signature) can be used to reproduce the process, provided that the referenced digital resource versions are available in data/ folder or elsewhere.
As part of the GloBI review process, such a DataBOM fingerprint (i.e., hash://md5/a9aaa7113716ab2f87ede6f6b70297c8) for the Afromoths dataset was published in [3] along with their referenced versioned resource. With this, you can re-generated the interpreted interaction records by running:
preston ls \
--algo md5 \
--remote https://zenodo.org \
--anchor hash://md5/a9aaa7113716ab2f87ede6f6b70297c8 \
| elton stream \
--algo md5 \
--data-dir data \
--prov-dir data \
--anchor hash://md5/a9aaa7113716ab2f87ede6f6b70297c8 \
| head -n2
where the DataBOM fingerprint hash://md5/a9aaa7113716ab2f87ede6f6b70297c8 is used to define (or anchor) the resources needed to produced the interpreted interaction data records. With this, we established a way to reproduce the process to generate interaction records without having the assume that ChecklistBank data sevices remain available and produce expected results. Instead, we rely on an archived copy (e.g., [3]) of the process and the versioned resources used. And, because digital fingerprints [5] are location independent and storage media agnostic, this archived copy can be stored in whatever digital storage media may be available. In this case, we used Zenodo to store a copy of the Afromoths archive.
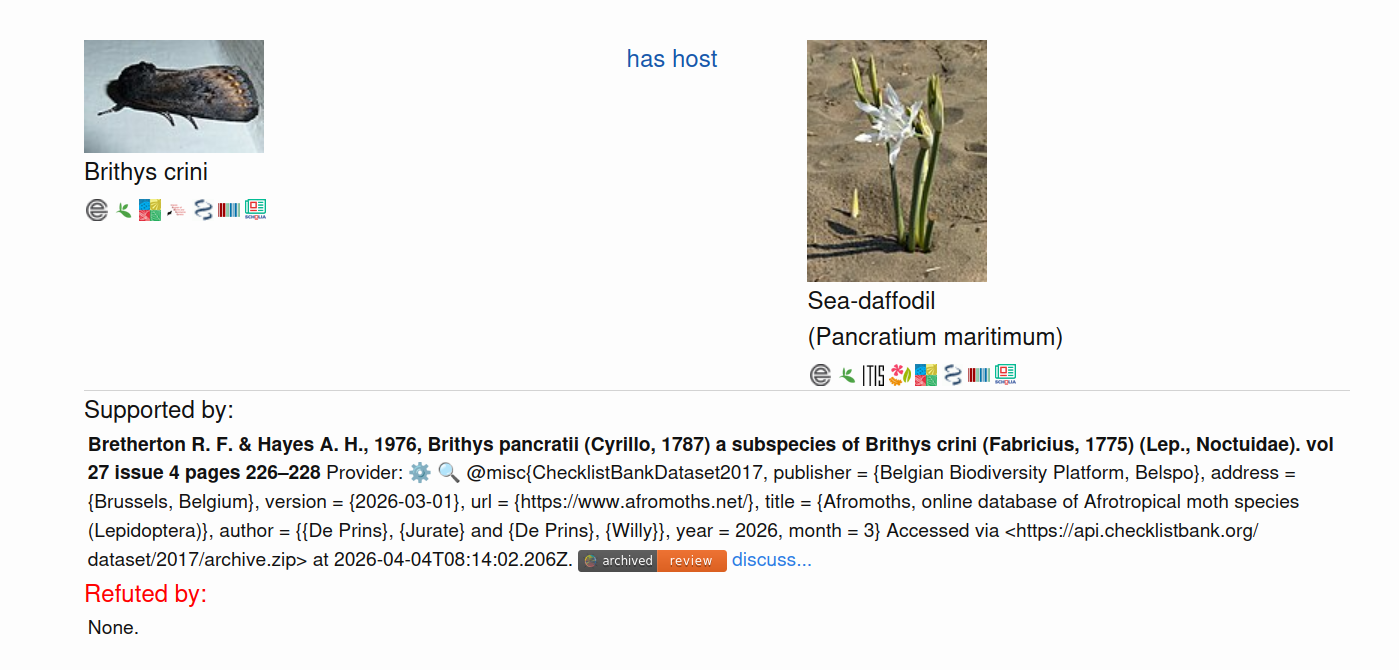
Additional reviews associated with the ChecklistBank datasets can be found via GloBI’s Zenodo Community or by clicking on the “archived review” badges on https://globalbioticinteractions.org/datasets or search results (see Figure 2.).
Discussion
Biodiversity data platforms such as ChecklistBank offer the ability to describe species interaction claims through registered datasets (e.g., [2], [6], [7]) using the Catalogue of Life Data Package (CoLDP) format. And, as of early 2026, GloBI helps discover these datasets via their search indexes, reviews and data products. This is yet another example how existing datasets in data infrastructures can be reused and integrated via review and data services associated with Global Biotic Interactions. With this, hopefully, these datasets can be more easily found just like their existing data cousins in the many other available data formats (e.g., DwC Associated Taxa, DwC Assocated Occurrences, DwC Resource Relations, or whatever other models happen to be in fashion).
A Note on CoLDP Schema Discovery and Dialects
On implementing GloBI’s support for extracting species interaction claims from ColDP formatted datasets, three distinct dialect were found. Because ColDP does not explicitly state the schema of the associated resources nor their location/name, elton implements ths following stategy to help interpret the available datasets:
- Query ChecklistBank for datasets with species interaction records.
- For each dataset, download their associated ColDP archive.
- Inspect each archive and attempt to find the species interaction table by scanning the archive for variations of filenames like
SpeciesInteractions.txt,SpeciesInteractions.csv,SpeciesInteractions.tsv,speciesinteractions.txt, …- Inspect the header of the species interaction table found, and attempt to detect one of three dialects used: (a) the TaxonWorks dialect (e.g.,
[6]) (b) the Belgium dialect (e.g.,[2]) and (c) the Hobern dialect (e.g.,[7]). See associated test cases and data package examples for more information.
Also, note that, unlike the relationshipOfResourceID of the DwC Resource Relationship Extension, the species interaction table in ColDP does not yet appear to support identifiers for the interaction type terms used. This is why a built-in translation table was introduced to map terms like “parasite of” to their RO equivalent http://purl.obolibrary.org/obo/RO_0002445 with label parasiteOf . Perhaps, a future version of the ColDP format will introduce support for species interaction type identifiers (e.g., typeID) to help more explicitly link the interaction type to their associated definition.
References
[1] Page, R.D.M. (2025). Using links from wikis to help discover content in the Biodiversity Heritage Library (BHL) Wikimedia and Biodiversity Data A Mutualistic Relationship in the Open Knowledge Ecosystem. Presented at Living Data/ Datos Vivos 2025 in Bogotá, Colombia on 2025-10-22. https://livingdata2025.com/program.html?abstract=7010913 https://youtu.be/wXaHJ85g88A?t=5814 .
[2] De Prins J., & De Prins W. (2006, October). Afromoths, online database of Afrotropical moth species (Lepidoptera). https://checklistbank.org/dataset/2017 https://doi.org/10.48580/d4fl.v124 https://www.afromoths.net/. Formerly published as a DwC archive through GBIF https://ipt.biodiversity.be/archive.do?r=afromoths https://www.gbif.org/dataset/65c9103f-2fbf-414b-9b0b-e47ca96c5df2 and https://doi.org/10.15468/s1kwuw .
[3] Elton, Nomer, & Preston. (2026). Versioned Archive and Review of Biotic Interactions and Taxon Names Found within urn:lsid:checklistbank.org:dataset:2017 hash://md5/a9aaa7113716ab2f87ede6f6b70297c8. Zenodo. https://doi.org/10.5281/zenodo.19389793.
[4] Döring, M., & Ower, G. (2019). The Catalogue of Life Data Package - A new format for exchanging nomenclatural and taxonomic information. Biodiversity Information Science and Standards, 3, e38771. https://doi.org/10.3897/biss.3.38771.
[5] Elliott M.J., Poelen, J.H. & Fortes, J.A.B. (2023) Signing data citations enables data verification and citation persistence. Sci Data. https://doi.org/10.1038/s41597-023-02230-y hash://sha256/f849c870565f608899f183ca261365dce9c9f1c5441b1c779e0db49df9c2a19d
[6] UCD Community. (2025). Universal Chalcidoidea Database curated in TaxonWorks (Apr 2025). https://doi.org/10.48580/d758p
[7] Hobern, D., & Gielis, C. (2025). Catalogue of the Pterophoroidea of the World (D. Hobern, Ed.; 1.1.25.304). Catalogue of Life, Amsterdam, Netherlands. https://doi.org/10.48580/d3gd.


